VyCAP’s single cell isolation solution is simple to use with high recoveries and results in high quality single cell DNA and RNA data. The system is designed to isolate rare single cells and compatible with all down-stream single-cell analysis technologies.
The single cell isolation solution comprises of two essential parts:
- A single-cell disposable, to distribute single cells in the wells of the isolation chip
- A Puncher system, to automatically select and isolate cells
VyCAP offers a complete validated and automated solution to isolate single cells. The solution comprises hardware, software and protocols. The work flows for single cells isolation a compatible with downstream single cell RNA or DNA analysis. Protocols to isolate single CTC and fetal cells are present on our website.
Unique Selling Points to isolate single cells
- Short workflow (<1hr) to obtain single cells,
- Large and small sample volumes 0.1-40m
- Automatically select and isolate cells
- Cell selection based on high quality immuno-fluorescence images
- Validated protocols for single cell DNA and RNA analysis
Isolation of single cells from whole blood for DNA and RNA analysis
The workflow to isolate cells from whole blood is validated and simple: it takes less than 1 hr to isolate rare cells such as CTC or fetal cells from whole blood. The single cell disposable contains a microwell chip with 6400 wells. Our single cell isolation system can be demonstrated in our demolab, contact us to learn if we can help.
One cell in one well – Chip with 6400 microwells available for isolation
VyCAP’s isolation chip consists out of 6400 microwells with a single pore in each well: the bottom of these wells are optically transparent with a thickness of 1µm. A cell suspension is transferred to the sample side of the disposable, containing this microwell, as the pump unit is activated. Hydrodynamic forces drag individual cells into the wells towards the pore in the bottom of the well. After a cell lands on the pore the sample flow through that well stops and no other cell will enter, diverting the next cell to a neighboring open well. In this manner single cells sort in individual wells across the entire chip; it typically takes 1-5 minutes to process a cell suspension and fill the microwells with single cells. To isolate single cells the well bottom together with a cell is punched into a PCR plate or PCR tube. Punching is fully automated and controlled by dedicated software.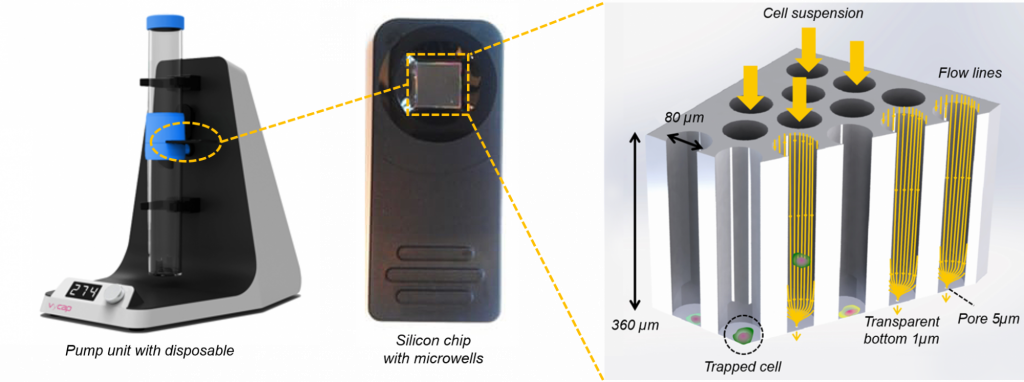
Puncher system is fully automated and easy to handle
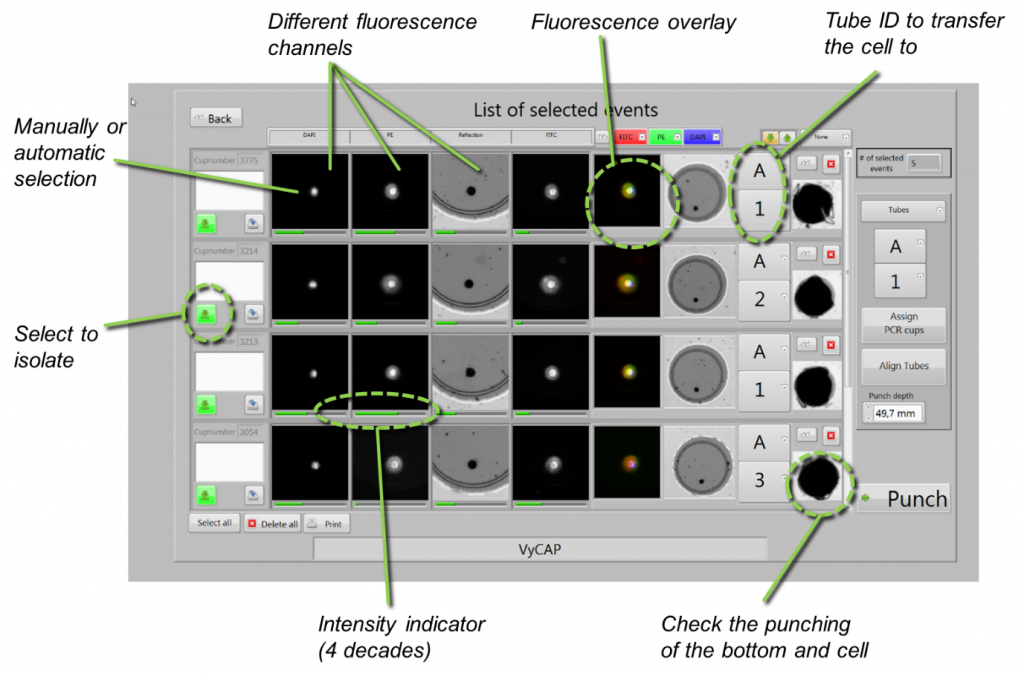
After filling of the microwells, the slide is transferred and easily loaded into the Puncher system to acquire fluorescent images. This system contains high end scanning stages to quickly position the chip, a Lumencor LED excitation light source and a Hamamatsu CMOS scientific camera. The stages are automatically controlled by the dedicated and user-friendly image acquisition software. Furthermore, the system has auto focus and an automatic filter cube changer that can hold a maximum of six different filter cubes. Below is an example of an image displaying a microwell chip filled with cells, stained with different fluorophores.
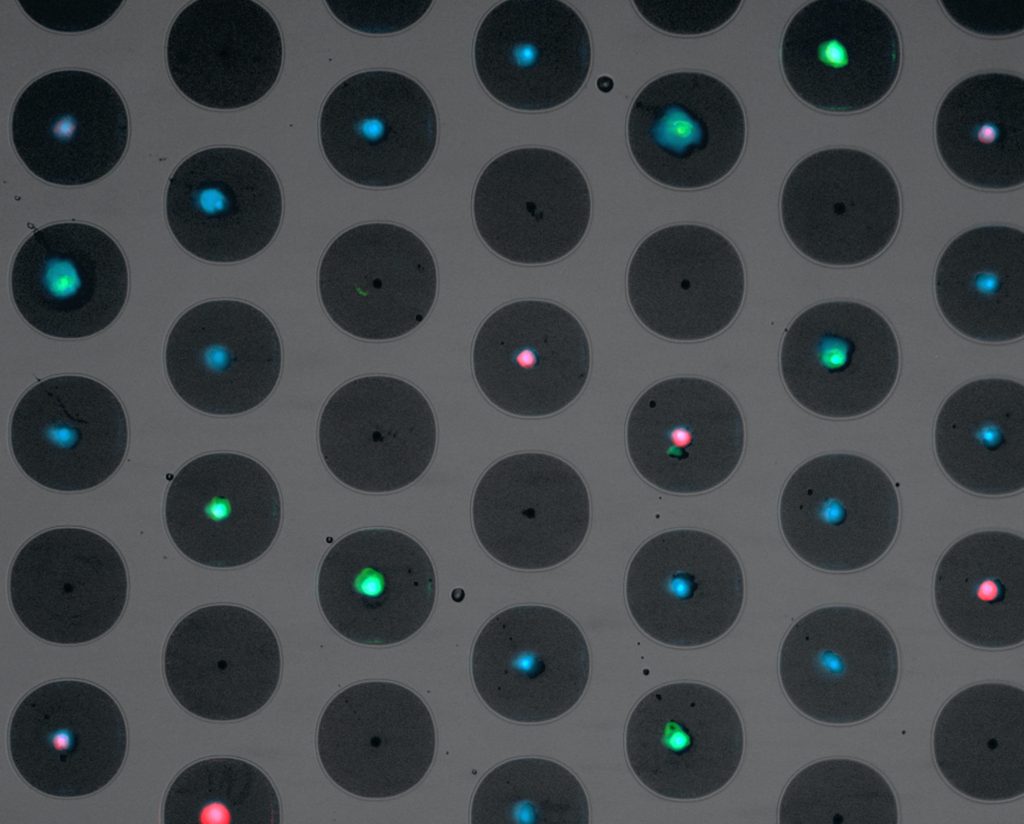

After image acquisition, the software can make a preselection of the cells of interest, based on the fluorescent cell signature. These preselected cells are presented in a gallery where cells can be selected for isolation by the signature Puncher technique.
Isolate single cells Punch by Punch
The selected cells are isolated using the innovative Puncher technology. The Puncher system automatically moves the slide towards the location of the punch needle after the desired container has been mounted into the device. The punch needle punches out the bottom of the well together with the cell in only a matter of a second. The alignment of the punch needle and the design of the tip of the punch needle is such that it does not touch the cell. The isolated cells are collected in a 0.2ml PCR tube or PCR plate. After punching an image of the punched microwell is acquired to ensure that the microwell bottom and cell are removed.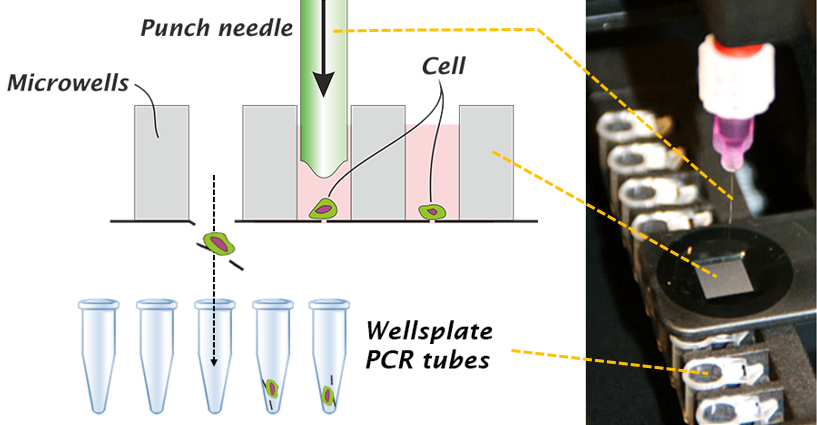
Validated WGA protocols for single cell DNA and RNA analysis
VyCAP has WGA protocols for viable and fixed cells using the Repli-G kit of Qiagen and the AMPLI-1 kit of Silicon Biosystems. The quality of the WGA product using these kits is determined by VyCAP’s WGA QC kit. The kit determines the presence of 10 DNA fragments present on 10 different chromosomes. For the REPLI-g kit all 10 bands are visible in the most optimal case. For the Ampli-1 kit, 7 bands are visible for maximum quality. One band is lost because the DNA fragment is too long to be amplified by the polymerase of the kit and two other fragments are lost due to the presence of restriction sites within these fragments, for the Mse-I enzyme as is used in the Ampli-1 WGA kit.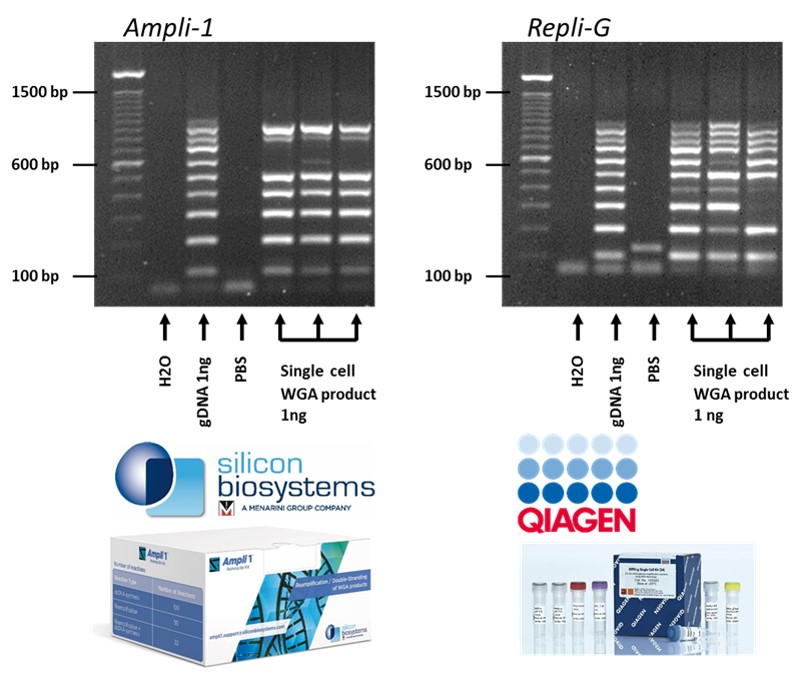
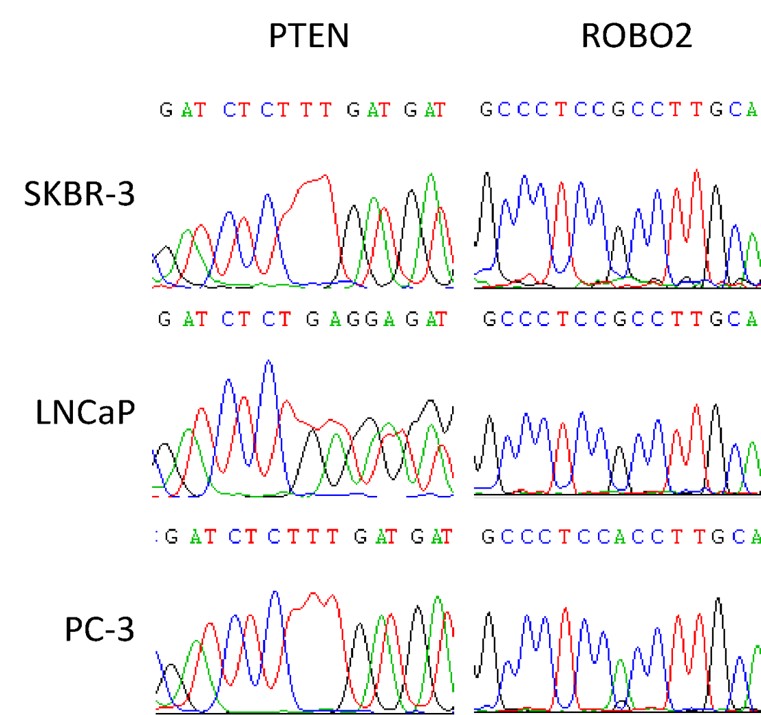
Single cell DNA analysis example
After single cell isolation and WGA the product is ready for DNA analysis. The example here displays the Sanger sequencing result for two genes PTEN and ROBO2. Three single cells of cell lines SKBR-3, PC-3 and LnCAP, are isolated using the Puncher platform technology, followed by AMPLI-1 WGA.